Publications
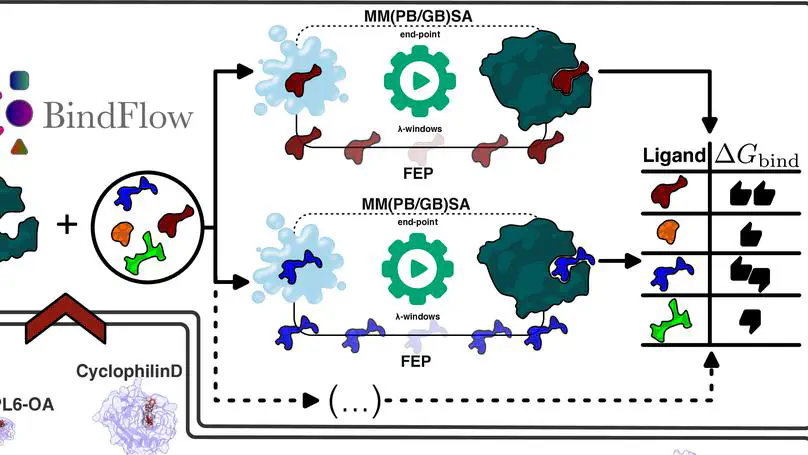
We present BindFlow, a Python-based software for automated absolute binding free energy (ABFE) calculations at the free energy perturbation (FEP) or at the molecular mechanics Poisson–Boltzmann/generalized Born surface area [MM(PB/GB)SA] level of theory. BindFlow is free, open-source, user-friendly, and easily customizable, runs on workstations or distributed computing platforms, and provides extensive documentation and tutorials. BindFlow uses GROMACS as a molecular dynamics engine and provides built-in support for the small-molecule force fields GAFF, OpenFF, and Espaloma, as well as support for user-provided custom force fields. We test BindFlow by computing affinities for 139 receptor–ligand pairs, involving eight different targets, including six soluble proteins, one membrane protein, and one nonprotein host–guest system. We find that the agreement of BindFlow predictions with experiments is overall similar to gold standards in the field. Interestingly, we find that MM(PB/GB)SA achieves correlations that, for some systems and force fields, approach those obtained with FEP while requiring only a fraction of the computational cost. This study establishes BindFlow as a validated and accessible tool for ABFE calculations.
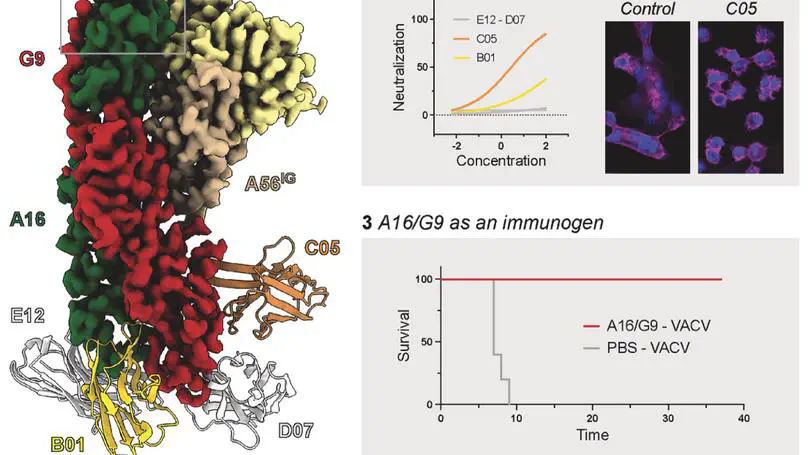
Monkeypox virus (MPXV) is a poxvirus endemic to Central and West Africa with high epidemic potential. Poxviruses enter host cells via a conserved entry-fusion complex (EFC), which mediates viral fusion to the cell membrane. The EFC is a promising therapeutic target, but the absence of structural data has limited the development of fusion-inhibiting treatments. Here, we investigated A16/G9, a subcomplex of the EFC that controls fusion timing. Using cryo-electron microscopy, we showed how A16/G9 interacts with A56/K2, a viral fusion suppressor that prevents superinfection. Immunization with A16/G9 elicited a protective immune response in mice. Using X-ray crystallography, we characterized two neutralizing antibodies and engineered a chimeric antibody that cross-neutralizes several poxviruses more efficiently than 7D11, the most potent antibody targeting the EFC described to date. These findings highlight the potential of A16/G9 as a candidate for subunit vaccines and identify regions of the EFC as targets for antiviral development.
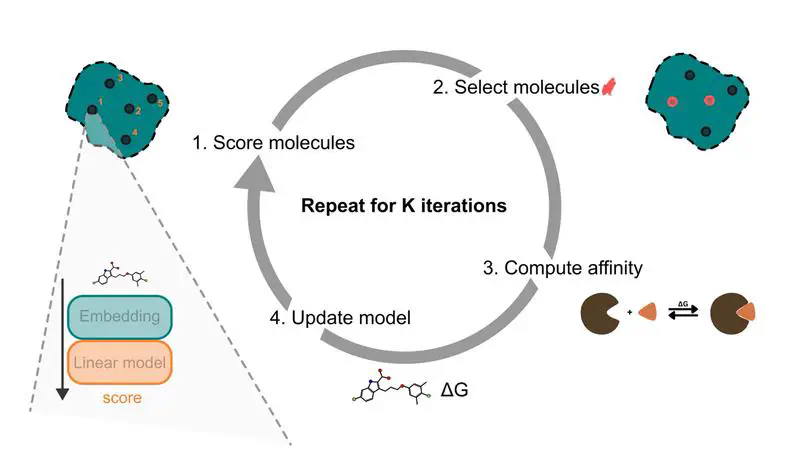
Predicting protein-ligand binding affinity with high accuracy is critical in structure-based drug discovery. While docking methods offer computational efficiency, they often lack the precision required for reliable affinity ranking. In contrast, molecular dynamics (MD)-based approaches such as MMGBSA provide more accurate binding free energy estimates but are computationally intensive, limiting their scalability. To address this trade-off, we introduce an active learning framework that automates molecule selection for docking and MD simulations, replacing manual expert-driven decisions with a data-efficient, model-guided strategy. Our approach integrates fixed - partly pre-trained deep learning - molecular embeddings (MolFormer, ChemBERTa-2, and Morgan fingerprints) with adaptive regression models (e.g. Bayesian Ridge and Random Forest) to iteratively improve binding affinity predictions. We evaluate this approach retrospectively on a new dataset of 60,000 chemically diverse compounds from ZINC-22 targeting the MCL1 protein using both AutoDock Vina and MMGBSA. Our results show that incorporating MMGBSA scores into the active learning loop significantly enhances performance, recovering 79.9% of the top 1% binders in the whole dataset, compared to only 6.7% when using docking scores alone. Notably, MMGBSA exhibits a stronger correlation with experimental binding affinities than AutoDock Vina on our dataset and enables more accurate ranking of candidate compounds in a runtime efficient way. Furthermore, we demonstrate that a one-at-a-time acquisition active learning strategy consistently outperforms traditional batched acquisition, the latter achieving just 78.4% recovery with MolFormer and Bayesian Ridge. These findings underscore the potential of integrating deep learning-based molecular representations with MD-level accuracy in an active learning framework, offering a scalable and efficient path to accelerate virtual screening and improve hit identification in drug discovery.

We present Moldrug, a computational tool for accelerating the hit-to-lead phase in structure-based drug design. Moldrug explores the chemical space using structural modifications suggested by the CReM library and by optimizing an adaptable fitness function with a genetic algorithm. Moldrug is complemented by Moldrug-Dashboard, a cross-platform and user-friendly graphical interface tailored for the analysis of Moldrug simulations. To illustrate Moldrug, we designed new potential inhibitors targeting the main protease (MPro) of SARS-CoV-2 by optimizing a consensus fitness function that balances binding affinity, drug-likeness, and synthetic accessibility. The designed molecules exhibited high chemical diversity. A subset of the designed molecules were ranked using MM/GBSA and alchemical binding free energy calculations, revealing predicted affinities as low as −10 kcal/mol. Moldrug is distributed as a Python package under the Apache 2.0 license. It offers pre-configured multi-parameter fitness functions for molecular design, while being highly adaptable for integrating functionalities from external software. Documentation and tutorials are available at https://moldrug.rtfd.io.
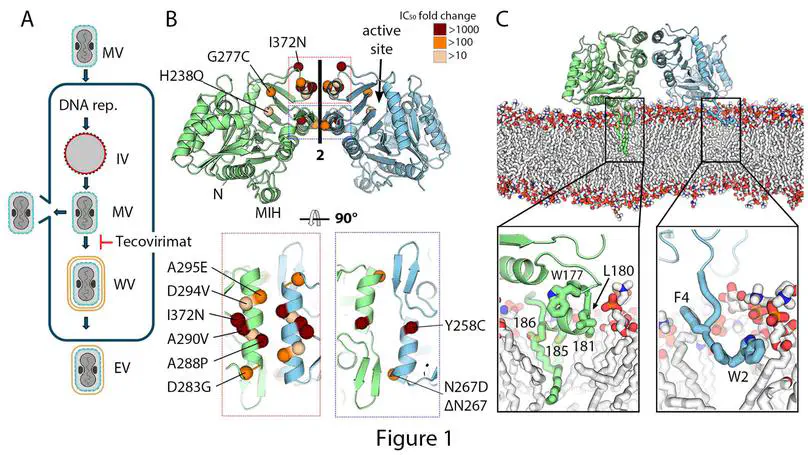
Mpox is a zoonotic disease endemic to Central and West Africa. Since 2022, two human-adapted monkeypox virus (MPXV) strains have caused large outbreaks outside these regions. Tecovirimat is the most widely used drug to treat mpox. It blocks viral egress by targeting the viral phospholipase F13; however, the structural details are unknown, and mutations in the F13 gene can result in resistance against tecovirimat, raising public health concerns. Here we report the structure of an F13 homodimer using X-ray crystallography, both alone (2.1 Å) and in complex with tecovirimat (2.6 Å). Combined with molecular dynamics simulations and dimerization assays, we show that tecovirimat acts as a molecular glue that promotes dimerization of the phospholipase. Tecovirimat resistance mutations identified in clinical MPXV isolates map to the F13 dimer interface and prevent drug-induced dimerization in solution and in cells. These findings explain how tecovirimat works, allow for better monitoring of resistant MPXV strains and pave the way for developing more potent and resilient therapeutics.